fatf.transparency.predictions.surrogate_explainers.TabularBlimeyLime¶
-
class
fatf.transparency.predictions.surrogate_explainers.TabularBlimeyLime(dataset: numpy.ndarray, predictive_model: object, as_regressor: bool = False, categorical_indices: Optional[List[Union[str, int]]] = None, class_names: Optional[List[str]] = None, feature_names: Optional[List[str]] = None)[source]¶ A tabular LIME explainer – a surrogate explainer based on a linear model.
Changed in version 0.1.0: (1) Added support for regression models. (2) Changed the feature selection mechanism from k-LASSO to
forward_selectionwhen the number of selected features is less than 7, andhighest_weightsotherwise – the default LIME behaviour.New in version 0.0.2.
This class implements Local Interpretable Model-agnostic Explanations (LIME) introduced by [RIBEIRO2016WHY]. This implementation mirrors the one in the official LIME package, which is available under the
lime.lime_tabular.LimeTabularExplainerclass therein.This explainer uses a quartile discretiser (
fatf.utils.data.discretisation.QuartileDiscretiser) and a normal sampler (fatf.utils.data.augmentation.NormalSampling) for augmenting the data. The following steps are taken to generate the explanation (when theexplain_instancemethod is called):The input
data_rowis discretised using the quartile discretiser. The numerical features are binned and the categorical ones are left unchanged (selected via thecategorical_indicesparameter).The data are sampled around the discretised
data_rowusing the normal sampler. Since after the discretisation all of the features are categorical the bin indices are sampled based on their frequency in (the discretised version of) thedatasetused to initialise this class.The sampled data are reverted back to their original domain and predicted with the black-box model (
predictive_modelused to initialise this class). This step is done via sampling each (numerical) feature value from the corresponding bin using the truncated normal distribution for which minimum (lower threshold), maximum (upper threshold), mean and standard deviation are computed empirically from all the data points from thedatasetfor which feature values fall into that bin. The categorical features are left unchanged.The discretised sampled data set is binarised by comparing each row with the user-specified
data_row(in theexplain_instancemethod). This step is performed by taking XNOR logical operation between the two – 1 if the feature value is the same in a row of the discretised data set and thedata_rowand 0 if it is different.The Euclidean distance between the binarised sampled data and binarised
data_rowis computed and passed through an exponential kernel (fatf.utils.kernels.exponential_kernel) to get similarity scores, which will be used as data point weights when reducing the number of features (see below) and training the linear regression.To limit the number of features in the explanation (if enabled by the user) we either use forward selection when the number of selected features is less than 7 or highest weights otherwise. (This is controlled by the
features_numberparameter in theexplain_instancemethod and by default –features_number=None– all of the feature are used.)A local (weighted) ridge regression (
sklearn.linear_model.Ridge) is fitted to the sampled and binarised data with the target being:The numerical predictions of the black-box model when the underlying model is a regression.
A vector of probabilities output by the black-box model for the selected class (one-vs-rest) when the underlying model is a probabilistic classifier. By default, one model is trained for all of the classes (
explained_class=Nonein theexplain_instancemethod), however the class to be explained can be specified by the user.
Note
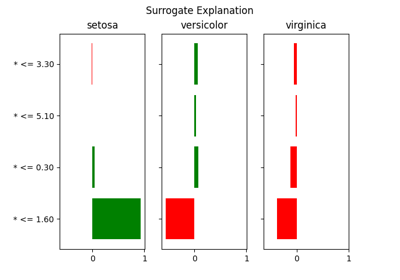
How to interpret the results?
Because the local surrogate model is trained on the binarised sampled data that is parsed through the XNOR operation, the parameters extracted from this model (feature importances) should be interpreted as an answer to the following question:
“Had this particular feature value of the explained data point been outside of this range (for numerical features) or had a different value (for categorical feature), how would that influence the probability of this point belonging to the explained class (probabilistic classification) / predicted numerical value (regression)?”
This LIME implementation is limited to black-box probabilistic classifiers and regressors (similarly to the official implementation). Therefore, the
predictive_modelmust have apredict_probamethod for probabilistic models andpredictmethod for regressors. When the surrogate is built for a probabilistic classifier, the local model will be trained using the one-vs-rest approach since the output of the global model is an array with probabilities of each class (the classes to be explained can be selected using theexplained_classparameter in theexplain_instancemethod). The column indices indicated as categorical features (via thecategorical_indicesparameter) will not be discretised.For detailed instructions on how to build a custom surrogate explainer (to avoid tinkering with this class) please see the How to build LIME yourself (bLIMEy) – Surrogate Tabular Explainers how-to guide.
For additional parameters, warnings and errors description please see the documentation of the parent class
fatf.transparency.predictions.surrogate_explainers.SurrogateTabularExplainer.- RIBEIRO2016WHY
Ribeiro, M.T., Singh, S. and Guestrin, C., 2016, August. Why should I trust you?: Explaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining (pp. 1135-1144). ACM.
- Parameters
- datasetnumpy.ndarray
A 2-dimensional numpy array with a dataset (utilised in various ways throughout the explainer).
- predictive_modelobject
A pre-trained (black-box) predictive model to be explained. If
as_probabilistic(see below) is set toTrue, it must have apredict_probamethod that takes a data set as the only required input parameter and returns a 2-dimensional numpy array with probabilities of belonging to each class. Otherwise, ifas_probabilisticis set toFalse, thepredictive_modelmust have apredictmethod that outputs a 1-dimensional array with (class) predictions.- as_regressorboolean, optional (default=False)
New in version 0.1.0.
A boolean indicating whether the global model should be treated as regression (
True) or probabilistic classification (False).- categorical_indicesList[column indices], optional (default=None)
A list of column indices in the input
datasetthat should be treated as categorical features.- class_namesList[string], optional (default=None)
A list of strings defining the names of classes. If the predictive model is probabilistic, the order of the class names should correspond to the order of columns output by the model. For other models the order should correspond to lexicographical ordering of all the possible outputs of this model. For example, if the model outputs
['a', 'c', '0']the class names should be given for['0', 'a', 'c']ordering.- feature_namesList[string], optional (default=None)
A list of strings defining the names of the
datasetfeatures. The order of the names should correspond to the order of features in thedataset.
- Attributes
- discretiserfatf.utils.data.discretisation.Discretiser
An instance of the quartile discretiser (
fatf.utils.data.discretisation.QuartileDiscretiser) initialised with the inputdatasetand used to discretise thedata_rowwhen theexplain_instancemethod is called.- augmenterfatf.utils.data.augmentation.Augmentation
An instance of the normal sampling augmenter (
fatf.utils.data.augmentation.NormalSampling) used to sample new data points around the discretiseddata_row(in theexplain_instancemethod).- bin_sampling_valuesDictionary[dataset column index, Dictionary[discretised bin id, Tuple(float, float, float, float)]]
A dictionary holding characteristics for each bin of each numerical feature. The characteristic are represented as a 4-tuple consisting of: the lower bin boundary, the upper bin boundary, the empirical mean of of all the values of this feature for data points (in
dataset) falling into that bin, and the empirical standard deviation (calculated in the same way). For the edge bins, if there are data available the lower edge is calculated empirically (as the minimum of the corresponding feature values falling into that bin), otherwise it is set to-numpy.inf. The same applies to the upper edge, which is either set tonumpy.infor calculated empirically (as the maximum of the corresponding feature values falling into that bin). If there are no data points to calculate the mean and standard deviation for a given bin, these two values are set tonumpy.nan. (This does not influence the future reverse sampling, for which this attribute is used: since there were no data for a given bin, the frequency of data for that bin is 0, therefore no data falling into this bin will be sampled.)
- Raises
- ImportError
The scikit-learn package is missing.
Methods
explain_instance(data_row, numpy.void], …)Explains the
data_rowwith linear regression feature importance.-
explain_instance(data_row: Union[numpy.ndarray, numpy.void], explained_class: Union[int, str, None] = None, samples_number: int = 50, features_number: Optional[int] = None, kernel_width: Optional[float] = None, return_models: bool = False) → Union[Dict[str, Dict[str, float]], Tuple[Dict[str, Dict[str, float]], Union[Dict[str, fatf.utils.models.models.Model], fatf.utils.models.models.Model]]][source]¶ Explains the
data_rowwith linear regression feature importance.Changed in version 0.1.0: Changed the feature selection mechanism from k-LASSO to
forward_selectionwhen the number of selected features is less than 7, andhighest_weightsotherwise – the default LIME behaviour.For probabilistic classifiers the explanations will be produced for all of the classes by default. This can be changed by selecting a specific class with the
explained_classparameter.The default
kernel_widthis computed as the square root of the number of features multiplied by 0.75. Also, by default, all of the (interpretable) features will be used to create an explanation, which can be limited by setting thefeatures_numberparameter. The data sampling around thedata_rowcan be customised by specifying the number of points to be generated (samples_number).By default, this method only returns feature importance, however by setting
return_modelstoTrue, it will also return the local linear surrogates for further analysis and processing done outside of this method.Note
The exact description of the explanation generation procedure can be found in the documentation of this class (
fatf.transparency.predictions.surrogate_explainers.TabularBlimeyLime).For additional parameters, warnings and errors please see the parent class method
fatf.transparency.predictions.surrogate_explainers.SurrogateTabularExplainer.explain_instance.- Parameters
- data_rowUnion[numpy.ndarray, numpy.void]
A data point to be explained (1-dimensional numpy array).
- explained_classUnion[integer, string], optional (default=None)
The class to be explained – only applicable to probabilistic classifiers. If
None, all of the classes will be explained. This can either be the index of the class (the column index of the probabilistic vector) or the class name (taken fromself.class_names).- samples_numberinteger, optional (default=50)
The number of data points sampled from the normal augmenter, which will be used to fit the local surrogate model.
- features_numberinteger, optional (default=None)
The maximum number of (interpretable) features – found with forward selection or highest weights – to be used in the explanation (the local surrogate model is trained with this feature subset). By default (
None), all of the (interpretable) features are used.- kernel_widthfloat, optional (default=None)
The width of the exponential kernel used when computing weights of the sampled data based on the distances between the sampled data and the
data_row.The defaultkernel_width(kernel_width=None) is computed as the square root of the number of features multiplied by 0.75.- return_modelsboolean, optional (default=False)
If
True, this method will return both the feature importance explanation dictionary and a dictionary holding the local models. Otherwise, only the first dictionary will be returned.
- Returns
- explanationsDictionary[string, Dictionary[string, float]]
A dictionary holding dictionaries that contain feature importance – where the feature names are taken from
self.feature_namesand the feature importances are extracted from local linear surrogates. These dictionaries are held under keys corresponding to class names (taken fromself.class_names).- modelssklearn.linear_model.base.LinearModel, optional
A dictionary holding locally fitted surrogate linear models held under class name keys (taken from
self.class_names). This dictionary is only returned when thereturn_modelsparameter is set toTrue.
- Raises
- TypeError
The
explained_classparameter is neitherNone, an integer or a string. Thesamples_numberparameter is not an integer. Thefeatures_numberparameter is neitherNonenor an integer. Thekernel_widthparameter is neitherNonenor a number. Thereturn_modelsparameter is not a boolean.- ValueError
The
samples_numberparameter is a non-positive integer (smaller than 1). Thefeatures_numberparameter is a non-positive integer (smaller than 1). Thekernel_widthparameter is a non-positive number (smaller or equal to 0). Theexplained_classspecified by the user could neither be recognised as one of the allowed class names (self.class_names) nor an index of a class name.